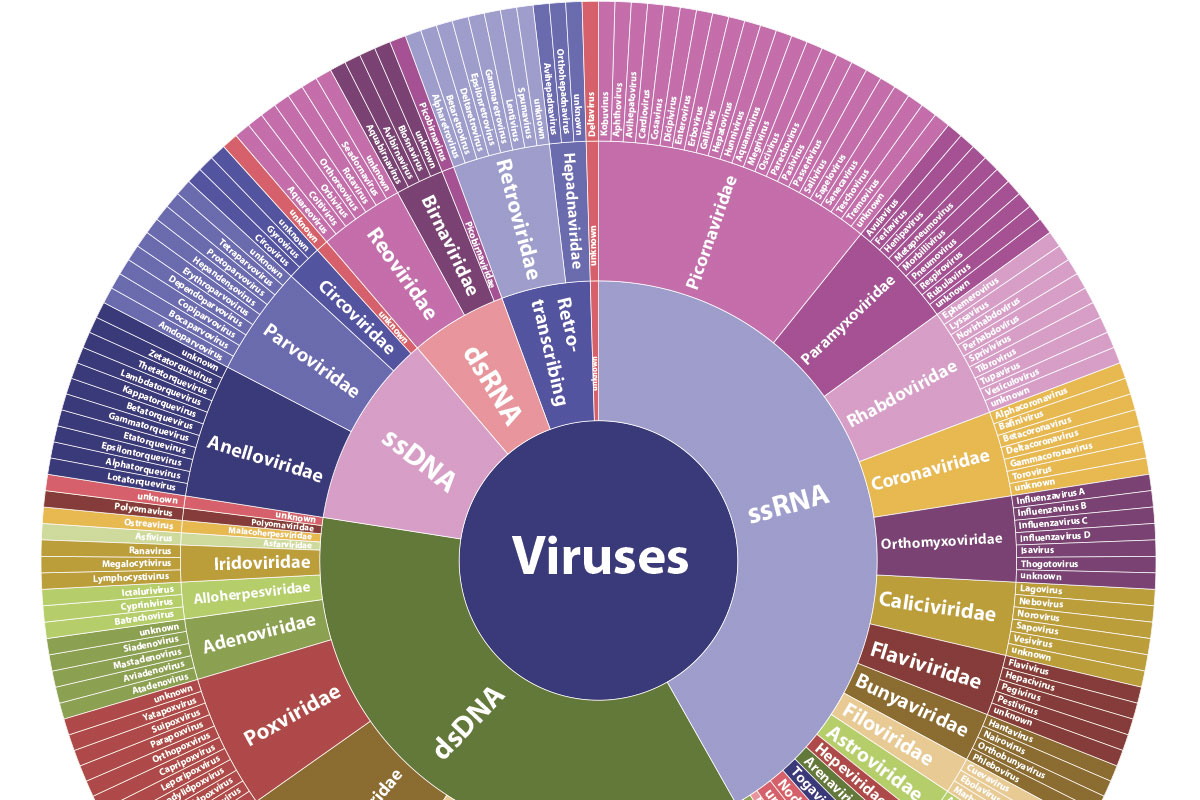

Thousands of viruses plague the human body, spawning everything from the mildly annoying common cold to deadly illness, and new threats continue to appear. When serious disease strikes, the question immediately becomes: “Which virus is it?”
Doctors must act on well-informed suspicions to order the necessary tests. And low levels of bad bugs can evade detection. Even after an exhaustive workup, the cause may remain a mystery.
A new test developed at Washington University School of Medicine is capable of detecting virtually any virus that infects
people and animals and could make such uncertainty a thing of the past.

“With this test, you don’t have to know what you’re looking for,” said Gregory A. Storch, MD, the study’s senior author and the Ruth L. Siteman Professor of Pediatrics. “It casts a broad net and can efficiently detect viruses that are present at very low levels. We think the test will be especially useful in situations where a diagnosis remains elusive after standard testing or in situations in which the cause of a disease outbreak is unknown.”
The test could be used to detect outbreaks of deadly viruses such as Ebola, Marburg and severe acute respiratory syndrome (SARS), as well as more routine viruses, including rotavirus and norovirus, both of which cause severe gastrointestinal infections. Storch’s team already used an earlier version of the test to determine which particular strain of the respiratory virus, enterovirus D-68, was causing an outbreak of unusually severe illness in children last year.
The Washington University team is making the technology publicly available for the benefit of patients and research.
Developed in collaboration with the university’s Elizabeth H. and James S. McDonnell III Genome Institute, the new test sequences and detects viruses in patient samples and is just as sensitive as the gold-standard polymerase chain reaction (PCR) assays, which are used widely in clinical laboratories. However, even the most expansive PCR assays can only screen for up to about 20 similar viruses at the same time.
Results published in the journal Genome Research demonstrate that the new test, called ViroCap, can detect viruses not found by standard testing of patient samples.

The researchers evaluated the new test by comparing it with an advanced genome sequencing technique called metagenomic shotgun sequencing (MSS). Storch and his colleagues were pioneers in using MSS to detect viruses. They performed both tests on sets of biological samples — including blood, stool and nasal secretions — from patients at St. Louis Children’s Hospital. The MSS test detected viruses in 10 of 14 patients. But the new ViroCap test found viruses in the four children that earlier testing had missed. MSS testing also failed to detect common, everyday viruses, including influenza B, a cause of seasonal flu; parechovirus, a mild gastrointestinal and respiratory virus; herpes virus 1, responsible for cold sores in the mouth; and varicella-zoster virus, which causes chickenpox.
In a second group of children with unexplained fevers, MSS testing had detected 11 viruses in the eight children evaluated. But the new ViroCap test found another seven, including a respiratory virus called human adenovirus B type 3A, which usually is harmless but can cause severe infections in some patients.
In all, the number of viruses detected in the two patient groups jumped to 32 from 21, a 52 percent increase.
“The test is so sensitive that it also detects variant strains of viruses that are closely related genetically,” said corresponding author Todd Wylie, an instructor of pediatrics. “Slight genetic variations among viruses often can’t be distinguished by currently available tests and complicate physicians’ ability to detect all variants with one test.”
Because the test includes detailed genetic information about various strains of particular viruses, subtypes can be identified easily. An example of this is the influenza virus, which may change yearly and has a variety of subtypes that could be responsible for a given outbreak. In the study, standard testing identified influenza A. The new test also could detect changes in the genome and indicated a particularly harsh subtype of the virus called H3N2.
Last flu season, H3N2 contributed to some 36,000 deaths in the U.S. For some patients, particularly young children, older adults and people with weakened immune systems, knowing that the H3N2 strain is present may alter treatment.
To develop the test, the researchers targeted unique stretches of DNA or RNA from every known group of viruses that infects humans and animals. In all, the research team included 2 million unique stretches of genetic material from viruses in the test. These stretches of material are used as probes that act like fishing bait to “catch” viruses in patient samples that are a genetic match. The matched viral material, which often includes most, if not all, of the virus’ genome, then is analyzed using high-throughput genetic sequencing. As novel viruses are discovered, their genetic material could be added to the test.
More research is needed to validate the test’s accuracy, so it could be several years before becoming clinically available. Already, the test is being used on patients enrolled in a clinical trial led by Storch.
“I think that in the future this test will provide information that will become very useful in clinical medicine far beyond what we get from an ordinary PCR test,” Storch said.
For now, scientists can use the technology to study viruses. Co-author Kristine Wylie, PhD, investigates the viruses that set up residence in and on the human body, collectively known as the virome. The test will provide a way to capture the full breadth and depth of such viruses and deepen understanding of how they play a role in keeping the body healthy.
“It also may be possible to modify the test so that it could be used to detect pathogens other than viruses, including bacteria, fungi and other microbes, as well as genes that would indicate the pathogen is resistant to treatment with antibiotics or other drugs,” said Kristine Wylie, PhD, an assistant professor of pediatrics.
“I think we’re going to see an explosion of research that grows out of this and it will have tremendous applications,” added Storch. “It will go far beyond what we’ve done.”
Published in the Winter 2015/2016 issue





 Share
Share Tweet
Tweet Email
Email