Announcing the Elizabeth H. and James S. McDonnell III Genome Institute
Genomics-based research is opening the door to more precise diagnoses, new disease treatments, and the ability to understand people’s health needs based on their genetic makeup. In the future, individualized DNA analysis will lead to a powerful form of preventive medicine.
To help support this paradigm shift in medicine, civic leaders and longtime philanthropists Elizabeth H. and James S. McDonnell III pledged $25 million to The Genome Institute, which will be renamed the Elizabeth H. and James S. McDonnell III Genome Institute at Washington University.
“This extraordinary gift will enable our top-line researchers to set more ambitious goals and step up discoveries in the fast-paced world of genomics,” said Washington University Chancellor Mark S. Wrighton.

Tracing the roots of disease
Under the leadership of Director Richard K. Wilson, PhD, the Alan A. and Edith L. Wolff Distinguished Professor of Medicine, and Co-director Elaine Mardis, PhD, the Robert E. and Louise F. Dunn Distinguished Professor of Medicine, the institute is helping to transform medicine using genomics.
Founded in 1993, the institute played a key role in the Human Genome Project, an international effort to decode all 6 billion letters of our genetic blueprint. It ultimately helped decode, or sequence, 25 percent of the genome by the project’s completion in 2003. The institute also has pioneered whole genome sequencing as a way to study cancer and other diseases.
In 2008, using private funds, institute researchers became the first to decode the whole genomes of both healthy and tumor cells from a leukemia patient — tracing the disease to its genetic roots. This potent mix of technology and ingenuity ushered in a new era, enabling researchers to search the entire genome to identify possible cancer-causing mutations.
Even though the institute is one of only three National Institutes of Health (NIH)-funded, large-scale sequencing centers in the U.S., it has struggled to find funding for some of its riskier projects. Sequencing the first whole cancer genome was a tough sell, as it was thought to be too expensive and too risky, possibly without much immediate benefit.

Family legacy
Elizabeth H. and James S. McDonnell III have championed research efforts at the School of Medicine for many years.
In 2000, together with Anne, AB ’64, and John F. McDonnell and the JSM Charitable Trust, they funded the construction of the McDonnell Pediatric Research Building, pictured above. The building was dedicated in memory of their daughter, Peggy, who died of cancer in 1972 at age 2.
In 2009, they endowed the Elizabeth H. and James S. McDonnell III Distinguished Professorship, which was instrumental in bringing Dennis Hallahan, MD, a preeminent figure in the study of cancers of the brain and central nervous system, to Washington University.
“Libby and Jim’s philanthropy has spawned scientific discoveries in nearly every pediatric discipline,” said Larry J. Shapiro, MD, executive vice chancellor and dean of the School of Medicine. “They have championed the medical school for many years, and this gift demonstrates their exceptional dedication to ensuring success in many key areas.”
A personal connection
The McDonnells, who lost their daughter, Peggy, at age 2 of neuroblastoma, a common cancer in children, have long supported pediatric cancer research. “What appeals to us about the institute is its collaboration with St. Louis Children’s Hospital, Washington University School of Medicine’s Department of Pediatrics and others in the application of genomics to pediatric cancers,” said James McDonnell III.
In 2010, to better understand the genetic origins of pediatric cancer, the institute launched the St. Jude Children’s Research Hospital-Washington University Pediatric Cancer Genome Project. The project has led to the whole genome sequencing of more than 1,000 childhood cancer patients, helping to pinpoint a variety of mutations, including a key mutation in pediatric neuroblastoma.
Over the past six years, cancer genomics studies at the institute have compared the tumor and normal genomes in patients with breast, liver, kidney, brain, lung, skin and blood cancers, among others.
“At some point, we’ll be able to look at many different patients with the same type of disease and start to understand what’s genetically similar about all these people’s diseases and what’s different,” Wilson said. “That information eventually will lead to the development of new treatments. But even in the short term we can use that information to more effectively utilize the drugs we already have on the shelf right now.”

Precision therapy
In one clinical study, Lukas Wartman, MD, a patient with leukemia, had relapsed twice and run out of options. Institute researchers sequenced his genome and discovered a mutation that might be treatable instead with a kidney cancer drug already on the market. Wartman, who is now an assistant professor in the Department of Medicine and an assistant director at the institute, took the drug and was able to attain remission, allowing him to receive a stem cell transplant. Without the sequencing data, Wartman’s doctors say he likely would not have survived.
“Soon, we’re going to see an era where patients are monitored much more closely, their diagnoses are much more precise and individualized and, hopefully, their outcomes are much better,” Mardis said. “Side effects will be diminished. The days of one-size-fits-all therapeutics will be gone.”
To fulfill the promise of putting genomics into clinical practice, Wilson said, philanthropy is crucial.
“There is not a lot of support at this time for clinical genomic testing of cancers because it’s still in its infancy,” he said. “To take it to the next level and really touch more patients, we’ve got to have gifts like the McDonnells bring.”

More data is better.
Solving a 6-billion-piece puzzle
Twins streamline the “big data” of genetics to aid clinical care
As sequencers churn out human genome data faster, and researchers better understand which “mistakes” in the genetic code lead to disease, a question remains: how to interpret and catalog the 6 billion letters of our genetic code so that clinicians can use it?
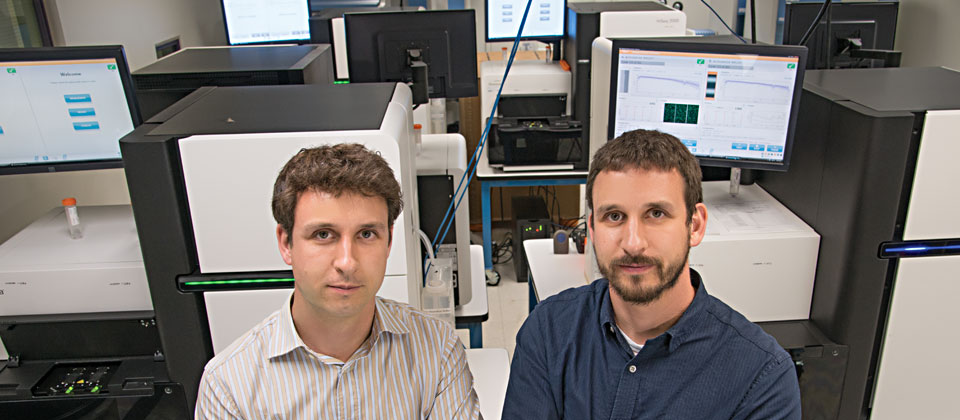
Obi Griffith, PhD, and Malachi Griffith, PhD, are identical twins and members of the McDonnell Genome Institute. They are streamlining information for clinicians and patients to aid in disease diagnosis and potential drug therapies. The brothers’ interest in pairing drugs with genetics is as much personal as it is scientific. Their mother died of breast cancer just weeks before they graduated from high school.
“The more data the better,” said Malachi Griffith, assistant professor of genetics in bioinformatics and associate director at the institute. “But our current capacity to deal with it means that it can often seem like too much to handle. The data is incredibly complex, and a lot of it is still unfamiliar to people practicing medicine.”
To date, the brothers and a team of software developers have launched two open-source websites, the Drug-Gene Interaction Database and the Database of Curated Mutations.
“With these tools, oncologists don’t have to read 100 papers,” said Obi Griffith, assistant professor of medicine and assistant director at the institute. “They can just get the final end product, the recommendation or the actionable statement.”