Humans are living in unprecedented times as multiple infectious diseases threaten to become public health issues. Ebola is grabbing headlines, but other important risks, such as Chikungunya and Enterovirus D-68, also have the potential to become the next big story.
“And, like death and taxes, there is always the flu,” said Steven J. Lawrence, MD, MSc, who works with health departments to prepare for, identify and respond to communicable, large-scale public threats.
“We don’t know how bad it will be, or what ages will be most affected each year, but we know it will be there.”
New and re-emerging infectious diseases pose a constant challenge. But whatever the threat, a network of physicians, researchers and other health-care professionals at Washington University Medical Center works determinedly to keep our community and the world safe from infectious diseases.
Preventing the next epidemic requires effort on many fronts: monitoring emerging diseases, conducting basic scientific research, training caregivers in specific patient-care protocols and advising public health officials on the appropriate response to outbreaks.
Scientists here are working to understand how viruses attack the body and evade the immune system and, ultimately, how to disable them. The Genome Institute at Washington University, one of only three National Institutes of Health (NIH)-funded, large-scale DNA sequencing centers in the U.S., frequently collaborates with researchers and physicians to pinpoint critical variations in pathogens, knowledge that can lay the foundation for disease treatments and discoveries.
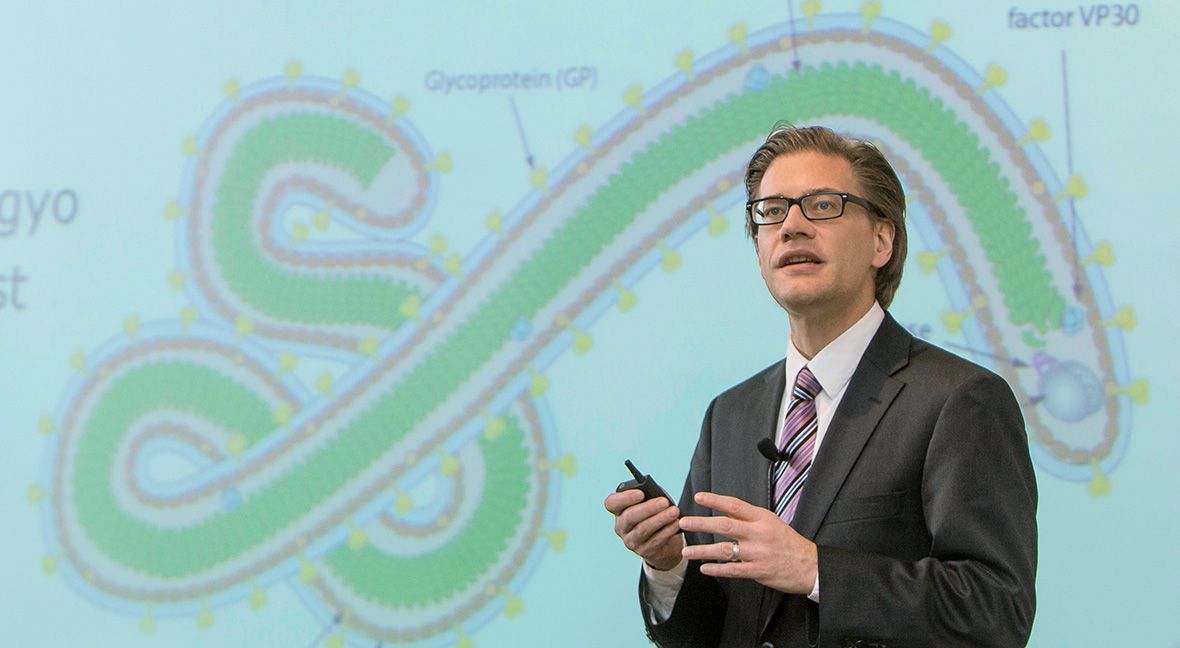

During a continuing medical education course, Steven J. Lawrence, MD, MSc, advises physicians on the safe management of Ebola patients. He and several other members of the infectious diseases faculty serve as spokespersons — responding to reporters when outbreaks occur and helping guide public response.
Borderless world
Many diseases — from anthrax to plague — begin in animals.
As people encroach on previously uninhabited spaces, there are more interactions with animals. Dozens, if not hundreds, of these naturally occuring infections in animals have yet to take root in humans, said Lawrence, an assistant professor of medicine in the Division of Infectious Diseases.
A case in point: The SARS coronavirus. This viral respiratory disease, recognized for the first time in humans in early 2003, originated in bats. The bats infected animals that later were sold for food in live markets in Guangdong Province, China. The disease spread to 29 countries — expedited through global airline travel — resulting in more than 8,000 cases and 776 fatalities.
Spearheaded by the World Health Organization, the international community responded with unheard-of speed, putting aside scientific rivalries and sharing data for the common good. Because of this, global data sharing is much more forthcoming today.
Washington University’s David Wang, PhD, then a postdoctoral student and now an associate professor of molecular microbiology and of pathology and immunology, gained international prominence when he and a small group of researchers used a new technology — a pan-viral microarray assay — to identify the novel SARS virus in 2003. Wang’s Pathogen Discovery and Infectious Disease Genomics Lab is a global player in identifying the guilty viral agent in disease outbreaks via advanced investigative technologies.
Amazingly, SARS has been eliminated in humans, purely through diagnostic testing and infection control, without the help of a vaccine or antiviral drug. Since the outbreak ended, there have been no new cases. “SARS had an almost 10 percent case fatality rate, and it was probably airborne in some instances,” Lawrence said. “It was really, really scary. Its elimination was a magnificent public health triumph.”
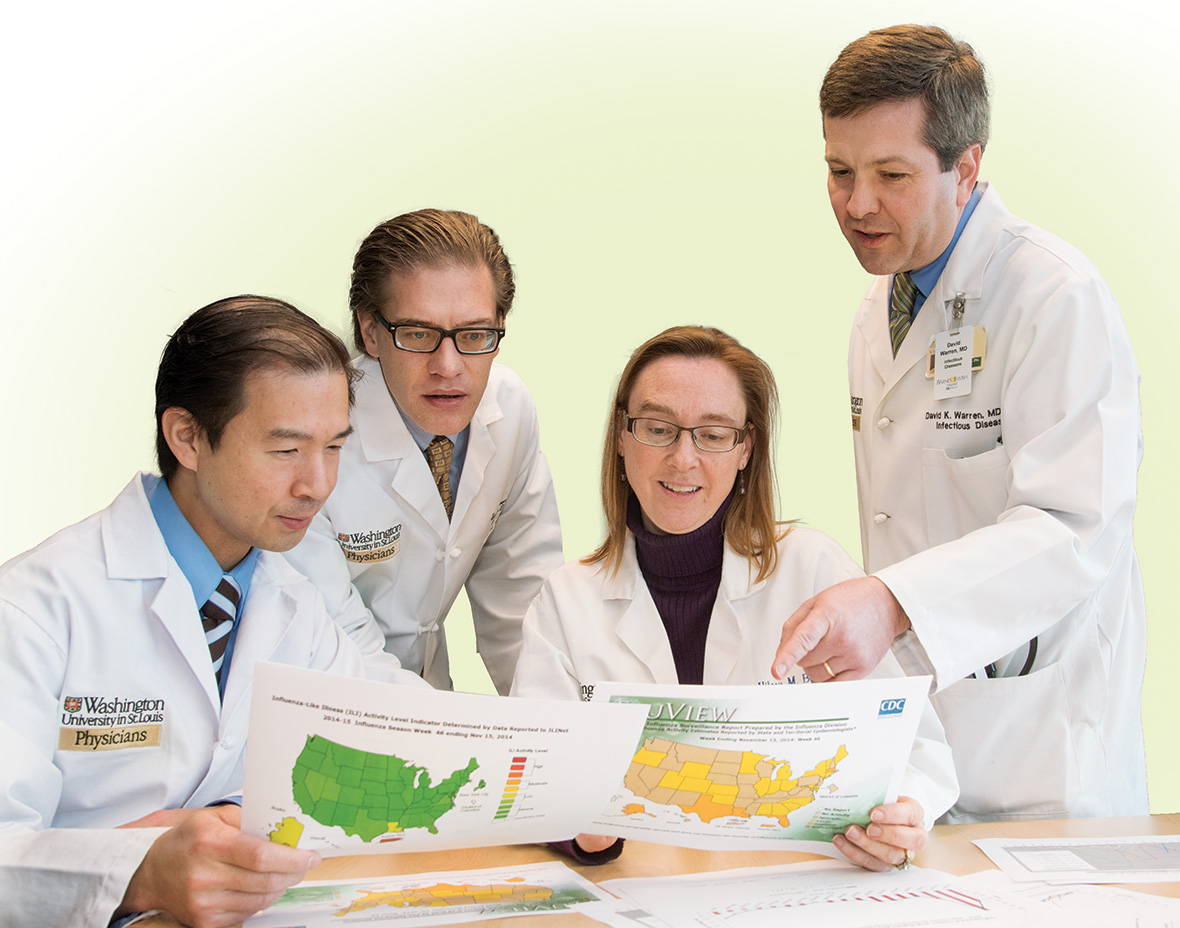

From left: Stephen Y. Liang, MD, medical director of Barnes-Jewish West County Infection Prevention and assistant professor; Lawrence; Hilary M. Babcock, MD, MPH, medical director, BJC Infection Prevention and Epidemiology Consortium, and associate professor; and David K. Warren, MD, MPH, hospital epidemiologist, Barnes-Jewish Hospital, and associate professor.
Disease detectives
In the early stages of an outbreak, physicians or public health officials often see something a little crazy, or outside the norm — a cluster of illnesses that test negative for the usual offenders.
In Queens, New York, in 1999, an astute doctor realized it was unusual for two cases of encephalitis to occur in a small neighborhood. Simultaneously, thousands of crows were dying in the city. West Nile Virus had just established itself in North America.
Early cases may go unnoticed until people start putting the puzzle together. Through tools such as ProMed-mail, an internet reporting system of the International Society for Infectious Diseases, physicians can log unusual symptoms. Dedicated to the rapid dissemination of information, the site includes official reports and accounts from media and local observers.
Washington University doctors closely follow the global situation and work hand in hand with the Centers for Disease Control and regional, state and local public health officials.
State laws require that health-care providers and laboratories report some types of infectious diseases to local health officials. St. Louis City Health Director Pamela Walker, MPH, regularly monitors results of statewide lab tests.
The health department follows up on all local reported disease cases, making sure infected people are receiving appropriate care and also tracing recent contacts to curb the spread. Guidance from Washington University doctors, Walker said, is critical in this process.
On a weekly basis, Walker seeks support from School of Medicine doctors on everything from sexually transmitted diseases to tuberculosis to health disparity issues. “I couldn’t possibly afford to pay for that level of expertise,” Walker said. “All of these physicians donate their time. The generosity is amazing. They just do it because they care.”
Lawrence, an epidemiologist, can explain infectious disease recognition and risk in terms that are easily understood, Walker said. Recently, he created a training video on Ebola, MERS (Middle East Respiratory Syndrome), Chikungunya and Enterovirus D-68 for health and human services staff, EMS workers and airport staff that every county in the region now uses.
As another means of surveillance, sensors in St. Louis can detect the release of deadly pathogens in the air in a terrorist attack. These sensors, also positioned in other major U.S. cities, are part of Homeland Security’s BioWatch program. Lawrence, who serves on the regional advisory committee for BioWatch, describes the sensors’ locations as “very sensitive information,” which terrorists could try to sabotage.
In October 2006, BioWatch sensors near Busch Stadium in downtown St. Louis received a positive signal for tularemia, or rabbit fever. This was a natural occurrence, not an act of terrorism. School of Medicine doctors helped health departments come up with good surveillance plans so that hospitals could identify and deal with cases of tularemia if any occurred.
Recognition is the important first step in containing any infectious disease threat, Lawrence said. “Once you recognize the illness, you can begin to determine the extent of it,” he said. “How many cases? Is it spread by direct contact? You can’t control it until you know how it is spread. The more you know about how a bug works, the more you can do to minimize the risk. You have to look at who’s gotten sick and who hasn’t. What are the risk factors?”
In the 1976 and 1995 Ebola outbreaks in the Democratic Republic of the Congo, it became clear that three groups were at a much higher risk: those who took care of an Ebola patient in their household or in a hospital and those involved in burial of Ebola patients. These same risk factors are playing out in today’s Ebola crisis.
While the population remains increasingly vulnerable to infectious diseases, technology is light-years ahead of where it was even 10 years ago. “A lot of this has to do with diagnostic testing and the ability to sequence new pathogens very quickly,” Lawrence said. “This in turn leads to the capacity to come up with possible treatments or vaccines much faster than in the past.”
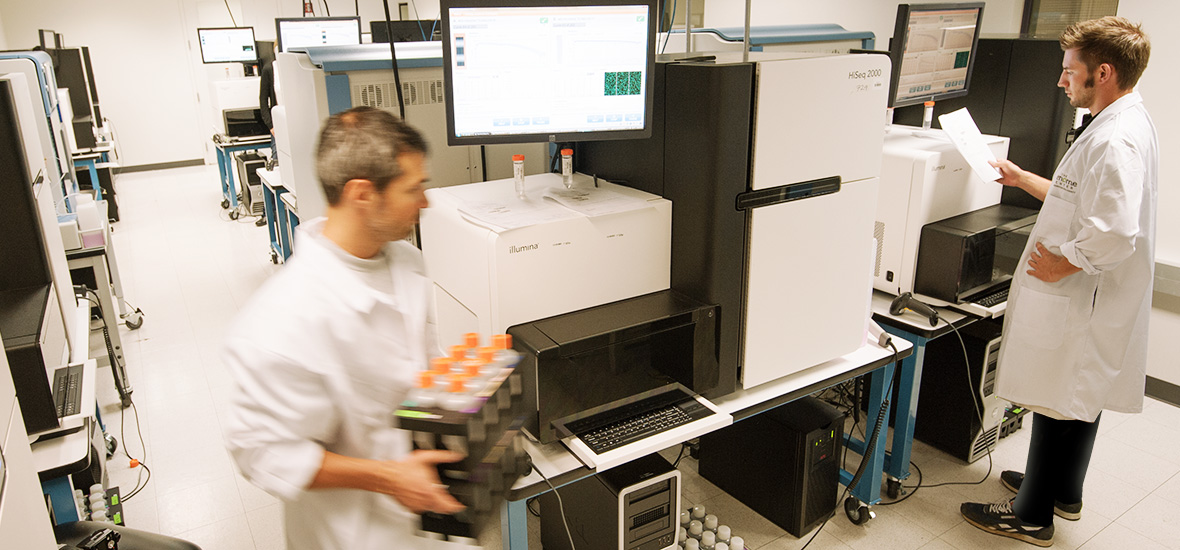

The program of investigation ranges from basic science inquiry to sequencing work at The Genome Institute.
Scientific discovery
In the Kansas City area last year, doctors began seeing severe respiratory illness in young children, with some requiring hospitalization or a ventilator. Nationwide, the virus, identified as Enterovirus D-68, spread rapidly; in rare cases, it has caused paralysis and death.
In collaboration with Greg Storch, MD, The Genome Institute recently sequenced the genome of Enterovirus D-68, sampled from patients treated at St. Louis Children’s Hospital. Kristine Wylie, PhD, research instructor in pediatrics, and Todd Wylie, director of microbial genomics computing, also were key members of the team. The results were immediately fed into the NIH genetic sequence database, GenBank, enabling access by scientists worldwide.
Storch, the Ruth L. Siteman Professor of Pediatrics, said the genetic data could be used to quickly develop a more reliable diagnostic test for the virus, so doctors will be able to confirm whether a child has the disease. There is no treatment and no vaccine, but having the DNA sequence available is a major step toward both goals.
Similarly, in October 2011, the St. Louis County Department of Health identified a cluster of E. coli-infected patients who had consumed salad bar items at supermarket chain stores. The outbreak sickened several dozen people in the Midwest, but not all cases had exposure to the suspected source.
Some questions could not be answered via lab methods routinely used in such investigations. Missouri State Epide-miologist George Turabelidze, MD, called in renowned expert on the bacterium, Phil Tarr, MD, the Melvin E. Carnahan Professor in Pediatrics and head of pediatric gastroenterology, who suggested using the resources of The Genome Institute. This marked the first time whole genome sequencing methodology was used for an E. coli outbreak investigation.
“Out of a couple million base pairs we found one or two nucleotide differences that sorted out epidemiologically perfectly,” Tarr said. “It validated the use of this technology to bring molecular clarity to outbreaks and focus public health resources. This work shows how powerful this technology can be. Now other people can build on these findings.”
The resulting study published by Tarr, Turabelidze and colleagues in the School of Medicine and at Washington State Public Health Laboratories, helped push the CDC to further speed up development of new technologies for outbreak investigations, Turabelidze said.
To protect patients and themselves, School of Medicine employees now must receive flu vaccinations per a 2013 policy change. Barnes-Jewish Hospital health-care professionals provide free flu shots to tens of thousands of St. Louisans annually.
Back to basics
Despite these advances, Lawrence said, time-tested medical know-how — contact tracing, hand hygiene and seasonal flu shots — can work wonders in stopping the spread of disease.
“It’s the simple things we’ve always known since John Snow (considered the father of epidemiology) traced a cholera outbreak to a water pump handle in 1854,” he said.
Unfortunately, previously controlled diseases are making a comeback due to lower vaccine compliance. The scientific community has done such a good job controlling diseases — most people today have never seen a case of polio, for instance — that many have become complacent.
Public concern is more focused on Ebola, but the chance of an uncontrolled Ebola outbreak in the U.S. is next to zero, Lawrence said. The disease is not easily transmitted, except in severely ill patients, and an advanced health-care infrastructure can effectively contain it.
“Old technology, basic infection control and isolation, stopped SARS,” he said. “It can also stop Ebola, even in the absence of widely available antiviral drugs to work against it.” While any case is tragic, Lawrence said, it’s important to note that of the more than 1,000 direct interactions between health-care workers and the nine Ebola patients on U.S. soil, there were two instances of transmission, as of December 2014.
“We can take care of an infection we’ve never seen in the U.S. before. We have a history of a strong infectious disease infrastructure to rapidly identify and put suspected patients in the right setting with the right kind of care — at Washington University Medical Center and in many U.S. hospitals,” Lawrence said. “The threats are always different, but there are always ways to defend against them.”
Published in the Winter 2014/2015 issue





 Share
Share Tweet
Tweet Email
Email